Ringer |
|
Documentation
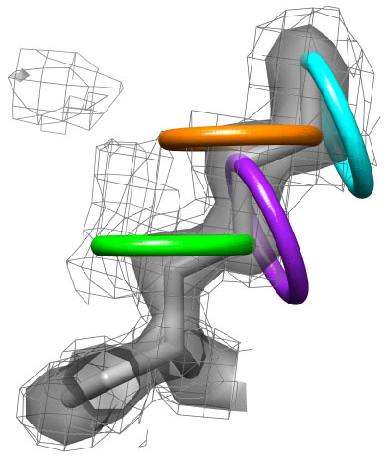
Ringer automatically samples electron density around the side-chain dihedral angles of a structural model and identifies peaks that may correlate with structural features. In contrast to manual methods, the program rapidly analyzes every residue in a model and applies consistent, objective criteria throughout the map to discover potential unmodeled side-chain ensembles. Peaks detected by Ringer currently require manual inspection and building of alternate conformers. Ringer is written in the programming language python and is distributed completely open source. It requires 20 MB of disk space to install. Ringer has undergone limited testing Debian Linux, Mac OS X, and Windows XP (using Cygwin) operating environments. Ringer is dependent on the external program Chimera, which is downloadable and freely available to academics. We also strongly recommend you also install either the CCP4 Software Suite or Phenix to generate electron density maps and the program Coot for interactively building structure models into electron density maps. When citing Ringer, please refer to: Lang PT and Ng H-L et al. Automated electron-density sampling reveals widespread conformational polymorphism in proteins. Protein Sci. 2010 Jul;19(7):1420-31. Older Versions
|