............... Stage must be at High Mag
............... move sample out out the beam by tapping on TouchScreen
............... Enter Command: "flux.com active"
............... move sample back by tapping "Undo" button on TouchScreen
To report FLux after the last "tuneup.com":
............... Enter Command: "flux.com last"
To report Flux after the last "tuneup.com" BEFORE an image taken at some previous time,
............... Enter Command: "flux.com /data/mcfuser/my_old_dir/image_name_x_xxxxx.cbf"
sample.com
usage: sample.com [x y z]
where: x is the distance to move in the direction of the x-ray beam y is the distance to move in the direction of gravity (down) z is the distance to move in the spindle direction (right-hand-rule)
or: sample.com to [x y z]
to implies that (x,y,z) is an absolute positon of the three sample motors x is the desired position of the first sample motor y is the desired position of the second sample motor z is the desired position of the third sample motor
example: sample.com returns: sample motors are now -0.623 -0.357 2.088
example: sample.com 0 0.1 0 returns: sample motors are now -0.722 -0.373 2.088
example: sample.com to -0.623 -0.357 2.088 returns: sample motors are now -0.623 -0.357 2.088Note in the second example that the relative move "down" by 100 microns changed both the x and y motors. This is because relative moves are done in the lab frame (x-ray beam, gravity, and spindle axis) and the motor moves are calculated with the current value of phi rotation in mind. This way, you can use the x coordinate to focus the sample in the mircoscope.
snap.com
usage: snap.com [filename.jpg] [camera] [352x240] [-mark] [-nodisplay]
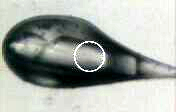
where: filename.jpg is the name of the image to safe [default: inherited from snapshot run] camera is the axis camera number to use [default: 1] 352x240 is the desired pixel dimensions of the image [default: 704x480] -mark will draw a circle indicating the beam profile on the image -nodisplay do not display the image after capturing it [default: display it] example: snap.com pretty.jpg -mark
tuneup.com is a full optimization of the x-ray beam through the pinhole. This procedure will move the sample 1mm down the cryostream (to protect it and to get it out of the way). Disable the feedback system. Insert the photodiode. Open the shutter. Maximize the flux out of the monochromator. Maximize the flux through the pinhole with a vertical and horizontal scan of the beam. Then the shutter is closed, diode removed and the feedback system recalibrated to the new beam position. Finally, the original sample position is restored. The hill-climbing features of the frontend software are used first, and then a "manual" scan through the current position is fitted to a parabola in gnuplot.
waypoint_setup.com
You must FIRST set up a "Run_1" in the bluice data collection tab before calling this script, then ... This script will create a copy-and-paste text (which you can then execute) to collect sets of images from different locations along your crystal; you will specify the starting and ending points AND the number of points. You can choose to set it up as a single continuous (Run_1) of frames (divided up and) collected at the different positions or, Muliple complete runs at different locations with identical parameters and incremented RUN numbers
Calculate the slit postition to achive a desired Attenuation of the Beam to reduce the Radiation Dose Rate
run "calculate_atten.com" from the command line
This calculates the horizontal divergence (divergence.com x) needed to obtain a given percentage
based on 100% full beam (2.0 milliradians; divergence.com 2.0)
.