MX Data Processing Workshop
Merged datasets:
goodsignal.mtz (815K)
badsignal.mtz (820K)
overlaps.mtz (804K)
decaying.mtz (824K)
icy.mtz (689K)
twin_5050.mtz (814K)
Electron density phased with the correct sites:
goodsignal.map (27M)
badsignal.map (27M)
overlaps.map (27M)
decaying.map (27M)
icy.map (27M)
twin_5050.map (27M)
The "right" model (used to calculate the data in the first place):
rightanswer.pdb (409K)
The "right" data sets (used to generate the images):
goodsignal_rightanswer.mtz (2.3M)
badsignal_rightanswer.mtz (2.3M)
Raw Image Data:
 goodsignal (2G)
goodsignal (2G)
or use wget -r bl831.als.lbl.gov/example_data_sets/Illuin/workshop/3dko/goodsignal/
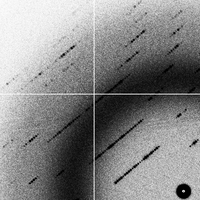 badsignal (2G)
badsignal (2G)
or use wget -r bl831.als.lbl.gov/example_data_sets/Illuin/workshop/3dko/badsignal/
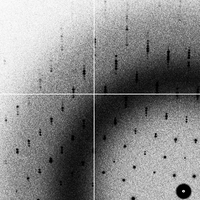 overlaps (2G)
overlaps (2G)
or use wget -r bl831.als.lbl.gov/example_data_sets/Illuin/workshop/3dko/overlaps/
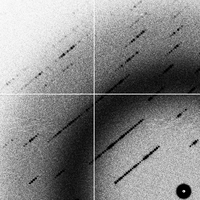 decaying (2G)
decaying (2G)
or use wget -r bl831.als.lbl.gov/example_data_sets/Illuin/workshop/3dko/decaying/
 icy (2G)
icy (2G)
or use wget -r bl831.als.lbl.gov/example_data_sets/Illuin/workshop/3dko/icy/
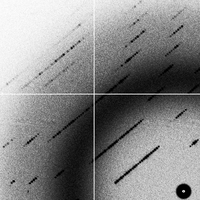 50-50 twin (2G)
50-50 twin (2G)
or use wget -r bl831.als.lbl.gov/example_data_sets/Illuin/workshop/3dko/twin_5050/
OR! Use the img_mix.com jiffy script to make a dataset with an
arbitrary twin fraction by making a weighted average of the "goodsignal" dataset above with
this "flipped" version of the "goodsignal" dataset:
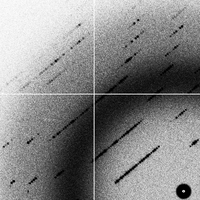 goodsignal_flipped (2G)
goodsignal_flipped (2G)
or use wget -r bl831.als.lbl.gov/example_data_sets/Illuin/workshop/3dko/goodsignal_flipped/
can you spot the differences between goodsignal and goodsignal_flipped?
You can also make yourself an arbitrary anomalous signal by mixing goodsignal with badsignal.
How low can you go and still solve it?
the mtz files are scaled and merged data processed with XDS. The maps are the results of mlphare/dm using the correct sites as run by Phaser Elves. The raw image data are also available. All data are derived from PDB id 3dko, but with Se atoms
substituted for the sulphur positions in that file.
Summary:
dataset completeness diff PADFPH map_CC R Rfree
goodsignal 99.97 7.5% 21.11 0.8104 8.84 11.99
badsignal 99.92 6.3% 4.38 0.0020 9.26 12.75
decaying 99.88 17.4% 17.52 0.5116 12.88 16.57
icy 91.58 62% 7.83 0.1033 72.83 71.30
overlaps 99.86 7.7% 20.44 0.8014 9.01 12.24
twin_5050 99.96 25% 12.61 0.3840 7.27 9.37
Description:
- The data set "goodsignal" corresponds to a perfectly adequate phasing signal with no overlap problems and no radiation damage.
- The "badsignal" data set is exactly the same, with the single exception that the native S atoms were NOT
replaced with Se. Notice how much harder it is to solve the structure?
- The "overlaps" data set is exactly the same as the "goodsignal" data set, but with the crystal orientation changed so that the c axis is nearly perpendicular to the rotation axis and overlaps are unavoidable.
- The "decaying" data set is also the same as the "goodsignal" data set, but half the sites are decaying exponentially from the beginning to the end and some non-isomorphism is creeping in.
- The "icy" data set is exactly the same as the "goodsignal" data set, except that it has ice rings.
- The "goodsignal_flipped" data set is also the same as the "goodsignal" data set, except it is in a different orientation.
- The "twin_5050" data set is a 50:50 mixture of the "goodsignal" and "goodsignal_flipped" images. The result of adding the images together in this way is identical to a crystal twinned with alpha=0.5.
-
"completeness" is the completeness of the final data set
-
"diff" is the difference between the final structure factors and the "right answer" (the structure factors used to generate the images).
-
"PADFPH" is the maximum peak height found in an anomalous difference Fourier map phased with the "right" phases, indicative of the anomalous signal strength.
- "map_CC" is the Pearson correlation coefficient of the "right answer" to the best possible experimental map (phased with absolutely correct heavy atom sites and solvent flattened).
- "R and Rfree" are the R factors after refining the "right answer" model against the structure factors obtained from these images.
 goodsignal (2G)
goodsignal (2G) badsignal (2G)
badsignal (2G) overlaps (2G)
overlaps (2G) decaying (2G)
decaying (2G) icy (2G)
icy (2G) 50-50 twin (2G)
50-50 twin (2G) goodsignal_flipped (2G)
goodsignal_flipped (2G)